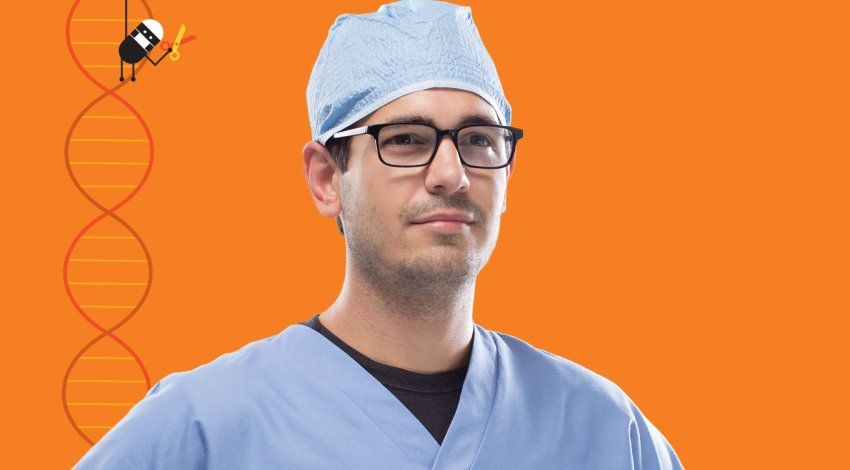
Scientists have discovered that the human body contains more than 25,000 genes, but what they do remains mostly a mystery. “We don’t know the function of the vast majority of genes,” says Nevan Krogan, PhD, director of the UC San Francisco branch of the California Institute for Quantitative Biosciences.
He and his team are seeking to crack this conundrum by examining genes and the proteins they encode through a superwide lens, as well as by creating large-scale physical and genetic interaction maps.
This approach turns traditional biological research on its head and is integral to precision medicine. In years past, most researchers studied genes or proteins one at a time, Krogan explains. That bottom-up pursuit usually began with a hypothesis and very little understanding of the enormously intricate structure housing the gene.
But once scientists had identified and mapped all the genes in the human genome, this extensive knowledge, along with sophisticated new tools, led to a top-down hunt. Investigators like Krogan now look globally at biological systems to better grasp the inner workings of cells. He’s particularly focused on the interactions between genes and the proteins they make, which perform a vast array of functions in the body.
“If you understand the machine in which the protein operates, then studying that protein, and the corresponding gene, becomes so much easier,” says Krogan, who is also an investigator at the UCSF-affiliated J. David Gladstone Institutes. Researchers gain this bird’s-eye view by analyzing a copious amount of data using complex computer algorithms.
If we can develop the tools to let the cells tell us their secrets without interjecting what we think we know, the data become so much more powerful.”
This unbiased strategy forgoes hypotheses and lets the data do the talking. “The cells are smarter than we are. If we can develop the tools to let the cells tell us their secrets without interjecting what we think we know, the data become so much more powerful,” he explains.
Krogan’s systems work is not only illuminating how genes and proteins function, it’s also shedding light on the underlying biology of disease for each person – a central goal of “omics” medicine, one of the pillars on which UCSF’s precision medicine platform rests.
Vexing Viruses
Infectious diseases caused by viruses are in Krogan’s crosshairs. Many viruses have fewer than two dozen genes but are a “fantastic tool for understanding molecular biology,” he says. “They basically hijack and rewire every major biological process in the cell they infect.”
To learn how viruses complete this assault, Krogan and his team tag viral proteins, inject them into a human cell, then pull them out. Next, using a tool called mass spectrometry, they identify which human (or host) proteins the viral proteins attack. “Essentially, we are taking a step back to look at the global landscape of what the virus needs during the course of infection,” he says.
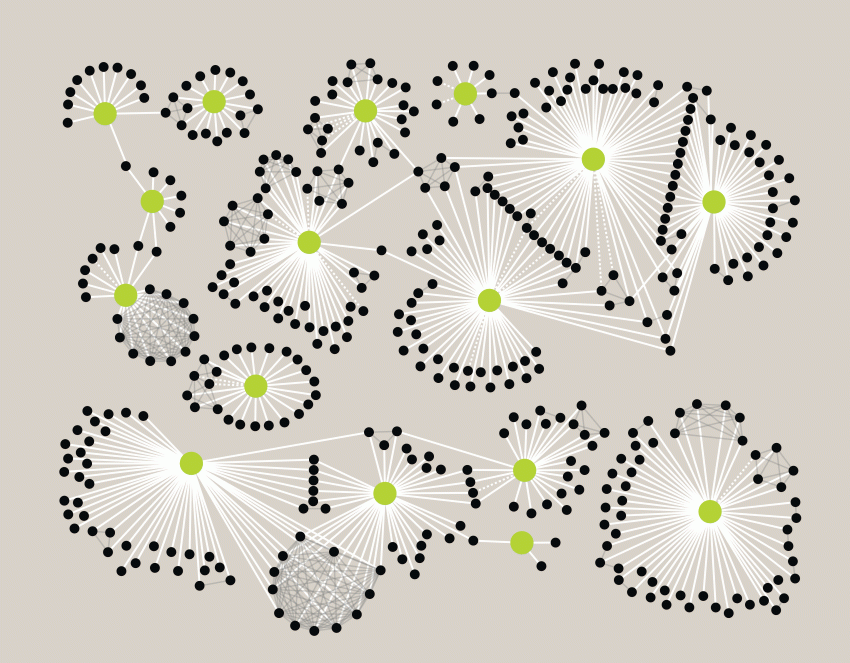
A protein-protein interaction network involving HIV and human proteins: HIV proteins (green nodes) and human proteins (black nodes) were found to physically interact (white lines). Interactions among human proteins (gray lines) reveal the molecular machines hijacked by the HIV proteins during the course of infection. Interrupting the connections among HIV and human proteins could provide future therapeutic strategies for the fight against AIDS. Illustration: Mike Shales
They started with HIV, injecting the 18 HIV proteins into a human cell. The team uncovered 400 human proteins hijacked by the pathogens. They subsequently analyzed a dozen other viruses, including polio, flu, herpes, hepatitis C, and dengue and discovered that the viral proteins commandeered a common set of host proteins.
These host proteins could serve as targets for new drugs, says Krogan. “If you can prevent these interactions, the cell has more of a fighting chance.” Directing drugs at protein interactions rather than at viruses could also avoid drug resistance, he notes, since viruses are able to mutate quickly and become resistant to drugs.
“Are these proteins ultimately going to serve as better therapeutic targets for future studies? I believe the answer is yes. We are trying for the silver bullet.”
Creating a Mega-Map to Master Mutations
Krogan is delving deeper into his examination of HIV by collaborating with Steven Wolinsky, MD, an infectious disease expert at Northwestern University. Wolinsky and several others have spent three decades monitoring a cohort of about 7,000 HIV-resistant men, trying to discover what makes them resistant. Krogan gave Wolinsky the list of 400 genes identified by his team as viral targets. Wolinsky is now sequencing these genes to determine if any mutations exist. If so, Krogan plans to apply his approach to study how these genes work to foster resistance.
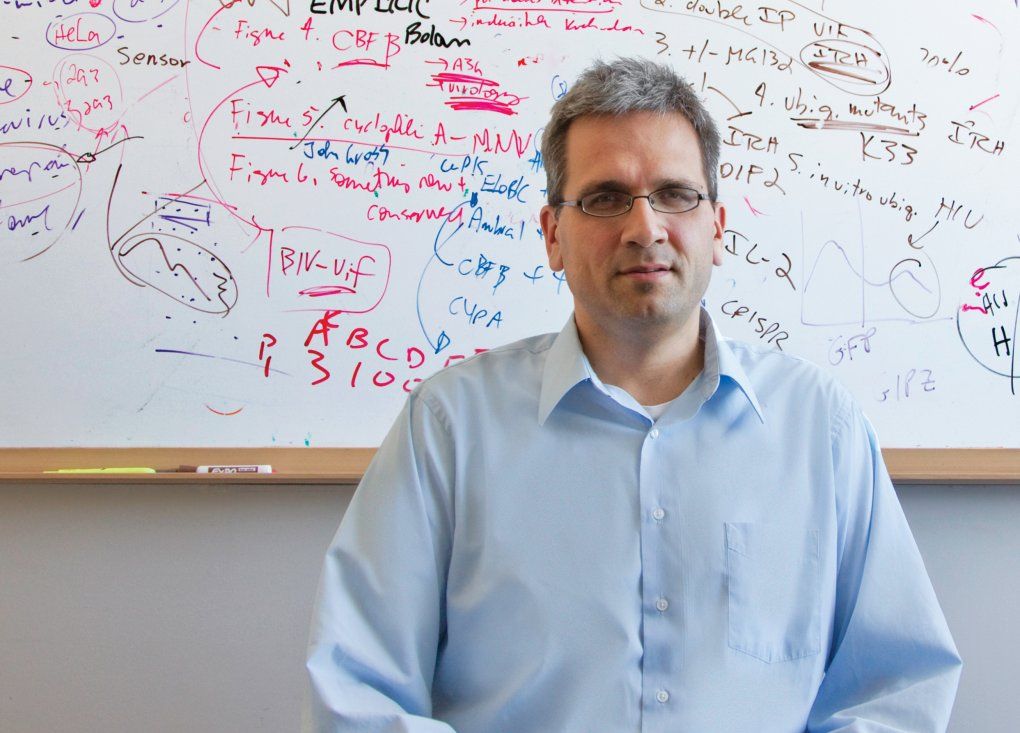

Molecular biologist Nevan Krogan is mapping how genes and proteins interact, knowledge that may lead to new drug targets. Photo: Susan Merrell
While he’s currently focusing on viruses, Krogan believes the same methodology can be used to reveal the underlying biology of other diseases. “Just like the genes in viruses conspire to produce infection, mutated genes work together in a devious fashion to create a disease state,” he says.
Science has made great strides in identifying sets of mutated genes that lead to certain diseases. “But if we don’t know what those genes do, how can we figure out how to fight them?” asks Krogan.
He wants to create large-scale maps of protein-protein and genetic interactions to provide a window into how all genes and proteins in a cell talk to one another – how they connect and interact both physically and functionally. It would be similar to a diagram of a machine showing how all the parts work together when the machine is running smoothly.
This comprehensive picture of cellular connectivity promises to help researchers pinpoint how unmutated genes function. Knowing that a protein is part of a four-protein complex, for example, would make studying that protein much simpler. Then a mutation could be introduced to see what unfolds. Are certain connections interrupted or pathways broken?
“Precision medicine could become more powerful because we would have a foundation with which to compare when genes go awry,” Krogan prophesies.